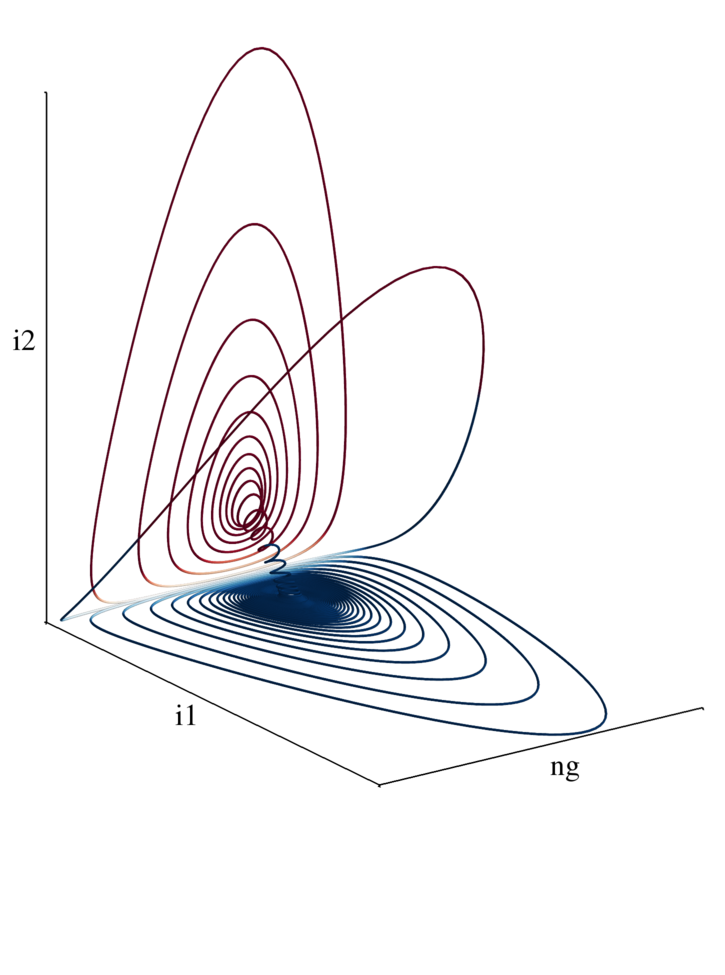
from mayavi import mlab import pandas as pd import numpy as np i1 = np.array(pd.read_csv('i1.dat')) i2 = np.array(pd.read_csv('i2.dat')) ng = np.array(pd.read_csv('ng.dat')) colors = np.zeros_like(i1) for i in xrange(colors.size): colors[i] = abs(i1[i] + i2[i]) if i1[i] < i2[i]: colors[i] = colors[i]*-1 mlab.options.offscreen = True fig = mlab.figure(size=(1800,2800),bgcolor=(1,1,1)) fig.scene.parallel_projection = True mlab.plot3d(20*ng, i1, i2,colors,colormap='RdBu',tube_radius=0.04,vmax=1.0,vmin=-1.0) ax = mlab.axes( color=(0,0,0)) ax.property.line_width = 3. ax.axes.label_format = '' ax.axes.x_label= '' ax.axes.y_label= '' ax.axes.z_label= '' mlab.draw() mlab.view( 55, 70, focalpoint='auto', distance='auto' ) mlab.savefig('mayavi_phase_space.png')
titles, axis lables, sizes etc
mostly inspired by this discussion.. here, Gael suggests using inkscape to add labels and stuff.. this is definitely a potential approach, however, if you'd like to be able to easily reproduce the plot in the future without having to mess around in a GUI, ImageMagick is a great choice..
so, to chop off the edges of the plot..
~/ $ convert mayavi_phase_space.png -gravity North -chop 0x600 -gravity South -chop 0x250 \
-gravity East -chop 200x0 -gravity West -chop 150x0 chopped_phase_space.png
and add our axis labels..
~/ $ convert chopped_phase_space.png -font Times-Roman -pointsize 60 -draw \
"text 25,700 'i2';text 350,1450 'i1';text 1100,1550 'ng'" annotated_phase_space.png
the nice thing about this approach is that it makes all messing about at the
end easily reproducible..
the finished product